
C57BL/6-Il22tm1(IL22)Bcgen Il22ra1tm1(IL22RA1)Bcgen/Bcgen • 121704

| Product name | B-hIL22/hIL22RA1 mice |
|---|---|
| Catalog number | 121704 |
| Strain name | C57BL/6-Il22tm1(IL22)Bcgen Il22ra1tm1(IL22RA1)Bcgen/Bcgen |
| Strain background | C57BL/6 |
| NCBI gene ID | 50616,58985 (Human) |
| Aliases | TIFa; IL-21; IL-22; ILTIF; IL-TIF; IL-D110; zcyto18; TIFIL-23; IL22R; CRF2-9; IL22R1 |
Gene targeting strategy for B-hIL22/hIL22RA1 mice.
The exons 1-5 of mouse Il22 gene that encode the whole molecule (ATG to STOP codon) are replaced by human counterparts in B-hIL22/hIL22RA1 mice. The promoter, 5’UTR and 3’UTR region of the mouse gene are retained. The human IL22 expression is driven by endogenous mouse Il22 promoter, while mouse Il22 gene transcription and translation will be disrupted. The exons 2-5 of mouse Il22ra1 gene that encode extracellular domain are replaced by human counterparts in B-hIL22/hIL22RA1 mice. The genomic region of mouse Il22ra1 gene that encodes transmembrane domain and cytoplasmic portion is retained. The promoter, 5’UTR and 3’UTR region of the mouse gene are also retained. The chimeric IL22RA1 expression is driven by endogenous mouse Il22ra1 promoter, while mouse Il22ra1 gene transcription and translation will be disrupted.
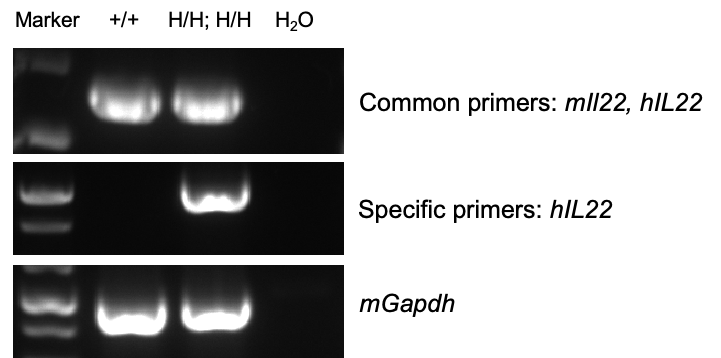
Species specific analysis of IL22 gene expression in wild-type C57BL/6 mice and homozygous humanized B-hIL22/hIL22RA1 mice by RT-PCR. Cerebellum were collected from wild-type C57BL/6 mice (+/+) and homozygous B-hIL22/hIL22RA1 mice (H/H; H/H). Common primers that detect both mIl22 and hIL22, as well as specific primers that only detect hIL22, were used in RT-PCR experiments. The positive bands obtained from common primers amplification were confirmed to be correct by sequencing. The specific primers for detecting hIL22 only detected hIL22 RNA in homozygous B-hIL22/hIL22RA1 mice. In summary, mouse Il22 mRNA was detectable in cerebellum from wild-type C57BL/6 mice. Human IL22 mRNA was detectable only in homozygous B-hIL22/hIL22RA1 mice, but not in wild-type C57BL/6 mice.
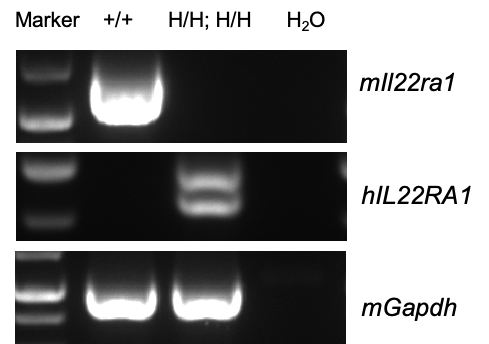
Species specific analysis of IL22RA1 gene expression in wild-type C57BL/6 mice and homozygous humanized B-hIL22/hIL22RA1 mice by RT-PCR. Small intestine were collected from wild-type C57BL/6 mice (+/+) and homozygous B-hIL22/hIL22RA1 mice (H/H; H/H). Mouse Il22ra1 mRNA was detectable only in wild-type C57BL/6 mice. Human IL22RA1 mRNA was detectable only in homozygous B-hIL22/hIL22RA1 mice, but not in wild-type C57BL/6 mice.
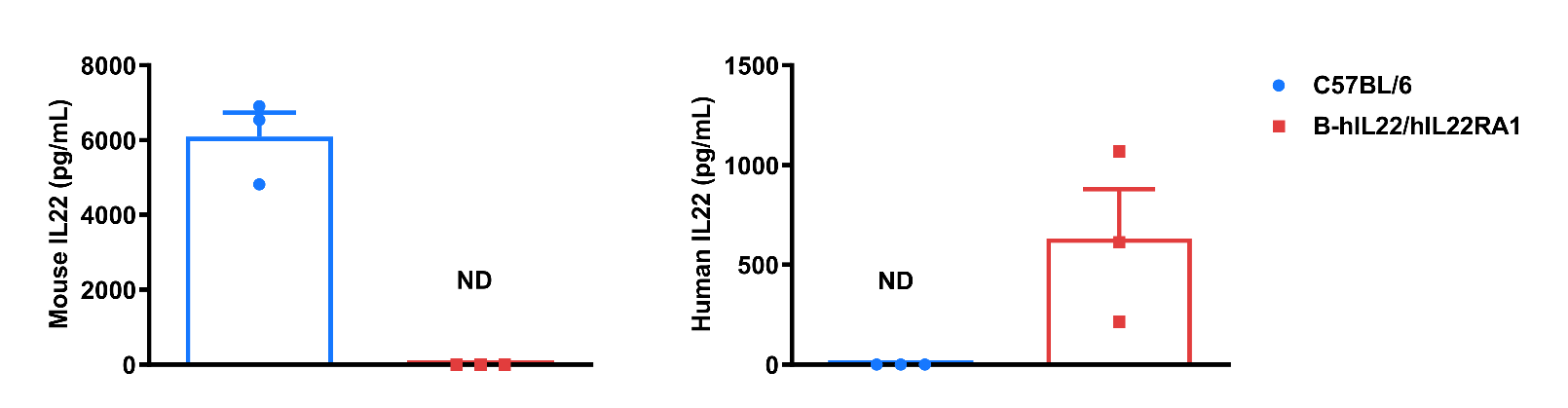
Strain specific IL22 expression analysis in wild-type C57BL/6 mice and homozygous humanized B-hIL22/hIL22RA1 mice by ELISA. Serum was collected from wild-type C57BL/6 mice (+/+) and homozygous B-hIL22/hIL22RA1 mice (H/H; H/H) stimulated with anti-mouse CD3ε antibody (7.5 μg/mouse, i.p.) and anti-mouse CD28 antibody (5 μg/mouse, i.p.) in vivo for 2 hrs (female, 14-week-old, n=3). Expression level of mouse and human IL22 were analyzed by ELISA (anti-mouse IL22 antibody: Abcam, ab223857; anti-human IL22 antibody: Abcam, ab216170). Mouse IL22 was only detectable in wild-type C57BL/6 mice. Human IL22 was exclusively detectable in homozygous B-hIL22/hIL22RA1 mice. Values are expressed as mean ± SEM. ND: not detectable.
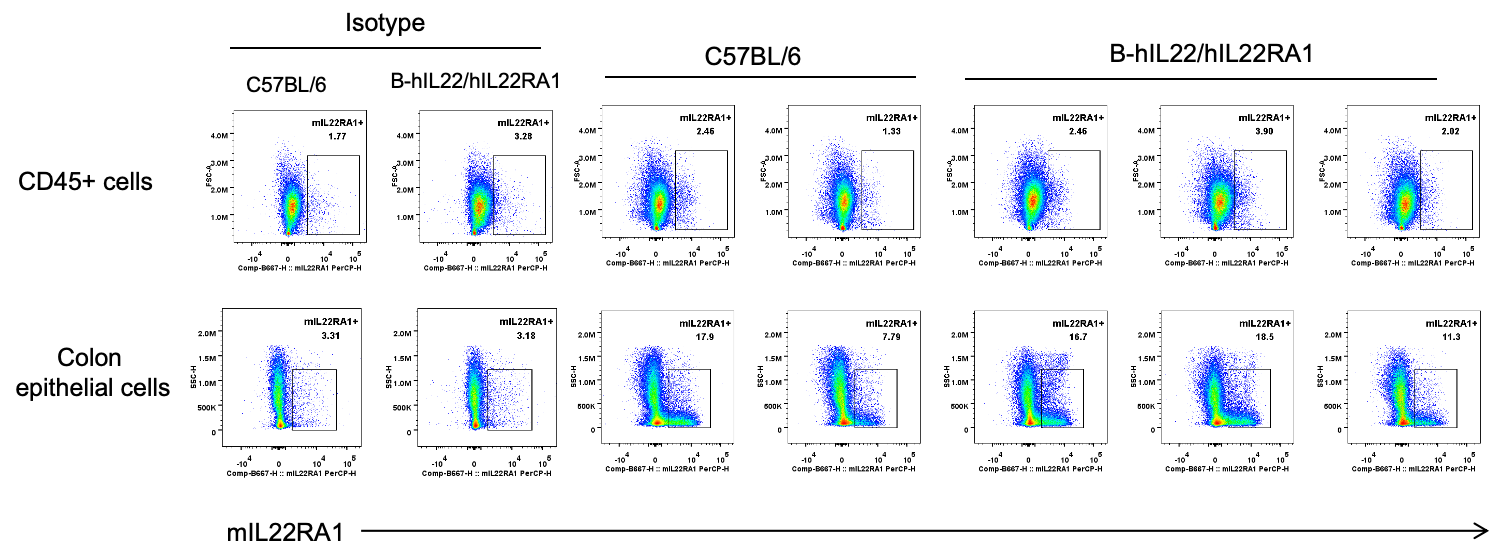
Strain specific IL22RA1 expression analysis in wild-type C57BL/6 mice and homozygous humanized B-hIL22/hIL22RA1 mice by flow cytometry. Colon epithelial cells were collected from wild-type C57BL/6 mice (+/+) and homozygous B-hIL22/hIL22RA1 mice (H/H; H/H) (female, 15-week-old, n=3). Protein expression was analyzed with anti-mouse IL22RA1 antibody (RD, FAB42941C) by flow cytometry. IL22RA1 was detectable in colon epithelial cells of wild-type C57BL/6 mice and homozygous B-hIL22/hIL22RA1 mice due to the cross-reactivity of antibodies, but not in CD45+ cells.
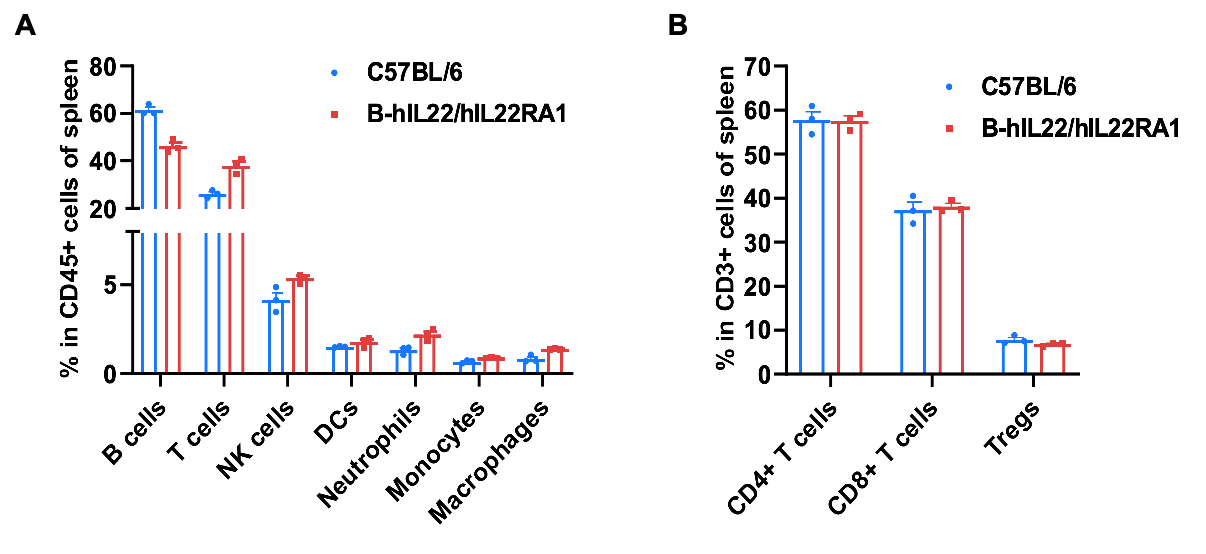
Frequency of leukocyte subpopulations in spleen by flow cytometry. Splenocytes were isolated from wild-type C57BL/6 mice and homozygous B-hIL22/hIL22RA1 mice (female, 6-week-old, n=3). A. Flow cytometry analysis of the splenocytes was performed to assess the frequency of leukocyte subpopulations. B. Frequencies of T cell subpopulations. Percentages of T cells, B cells, NK cells, DCs, monocytes, macrophages, neutrophils, CD4+ T cells, CD8+ T cells and Tregs in B-hIL22/hIL22RA1 mice were similar to those in C57BL/6 mice, demonstrating that humanization of IL22 and hIL22RA1 do not change the frequency or distribution of these cell types in spleen. Values are expressed as mean ± SEM.
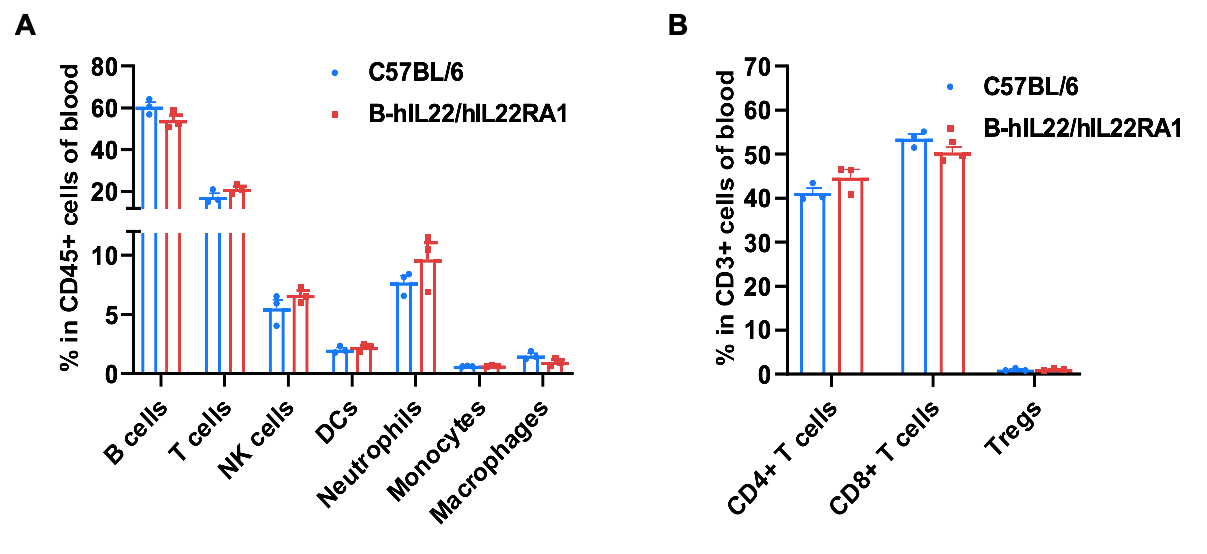
Frequency of leukocyte subpopulations in blood by flow cytometry. Blood cells were isolated from wild-type C57BL/6 mice and homozygous B-hIL22/hIL22RA1 mice (female, 6-week-old, n=3). A. Flow cytometry analysis of the blood cells was performed to assess the frequency of leukocyte subpopulations. B. Frequencies of T cell subpopulations. Percentages of T cells, B cells, NK cells, DCs, monocytes, macrophages, neutrophils, CD4+ T cells, CD8+ T cells and Tregs in B-hIL22/hIL22RA1 mice were similar to those in C57BL/6 mice, demonstrating that humanization of IL22 and IL22RA1 do not change the frequency or distribution of these cell types in blood. Values are expressed as mean ± SEM.
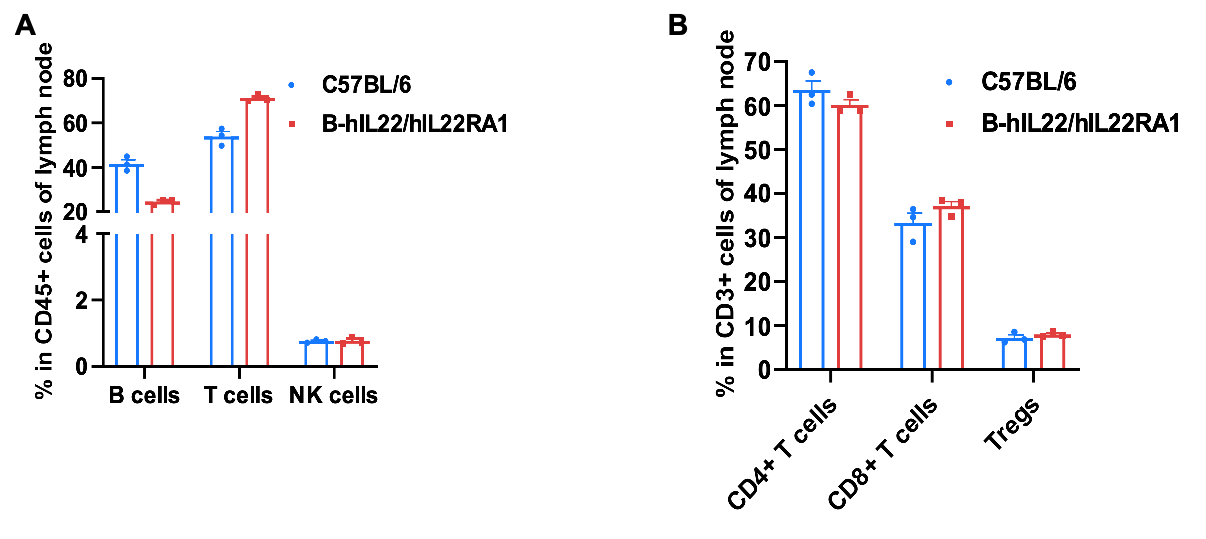
Frequency of leukocyte subpopulations in lymph node by flow cytometry. Leukocytes were isolated from wild-type C57BL/6 mice and homozygous B-hIL22/hIL22RA1 mice (female, 6-week-old, n=3). A. Flow cytometry analysis of the leukocytes was performed to assess the frequency of leukocyte subpopulations. B. Frequencies of T cell subpopulations. Percentages of T cells, B cells, NK cells, CD4+ T cells, CD8+ T cells and Tregs in B-hIL22/hIL22RA1 mice were similar to those in C57BL/6 mice, demonstrating that humanization of IL22 and IL22RA1 do not change the frequency or distribution of these cell types in lymph node. Values are expressed as mean ± SEM.